本周主要针对代码和数据进行了学习

主要先是把GEARS中用到的这些数据下载了下来并查看了数据的格式。
数据
Gene2Go
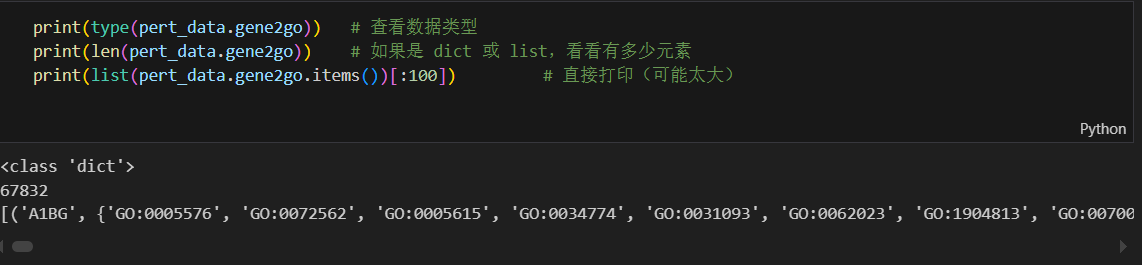
查看了Go数据集是怎么使用的,以及怎么检索GO数据的实际含义:
搜索GO的信息:https://geneontology.org/
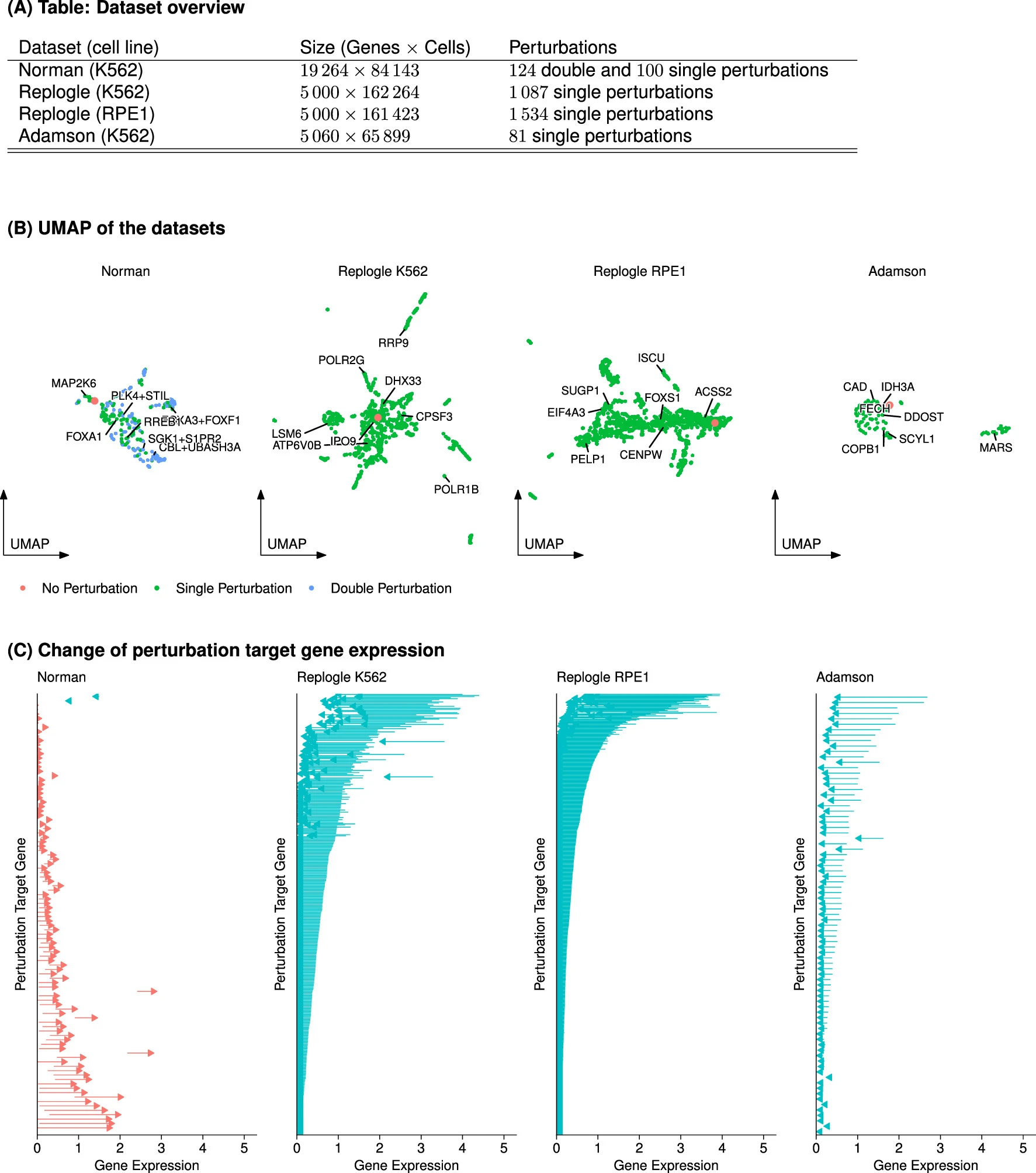
对于Norman数据集
print(adata_temp)
AnnData object with n_obs × n_vars = 91205 × 5045
obs: 'condition', 'cell_type', 'dose_val', 'control', 'condition_name'
var: 'gene_name'
uns: 'non_dropout_gene_idx', 'non_zeros_gene_idx', 'rank_genes_groups_cov_all', 'top_non_dropout_de_20', 'top_non_zero_de_20'
layers: 'counts'
print(adata_temp.obs['condition'])
cell_barcode
AAACCTGAGGCATGTG-1 TSC22D1+ctrl
AAACCTGAGGCCCTTG-1 KLF1+MAP2K6
AAACCTGCACGAAGCA-1 ctrl
AAACCTGCAGACGTAG-1 CEBPE+RUNX1T1
AAACCTGCAGCCTTGG-1 MAML2+ctrl
...
TTTGTCAGTCATGCAT-8 RHOXF2BB+SET
TTTGTCATCAGTACGT-8 FOXA3+ctrl
TTTGTCATCCACTCCA-8 CELF2+ctrl
TTTGTCATCCCAACGG-8 BCORL1+ctrl
TTTGTCATCTGGCGAC-8 MAP4K3+ctrl
Name: condition, Length: 91205, dtype: category
Categories (284, object): ['AHR+FEV', 'AHR+KLF1', 'AHR+ctrl', 'ARID1A+ctrl', ..., 'ZC3HAV1+HOXC13', 'ZC3HAV1+ctrl', 'ZNF318+FOXL2', 'ZNF318+ctrl']
print(adata_temp.obs['cell_type'])
cell_barcode
AAACCTGAGGCATGTG-1 A549
AAACCTGAGGCCCTTG-1 A549
AAACCTGCACGAAGCA-1 A549
AAACCTGCAGACGTAG-1 A549
AAACCTGCAGCCTTGG-1 A549
...
TTTGTCAGTCATGCAT-8 A549
TTTGTCATCAGTACGT-8 A549
TTTGTCATCCACTCCA-8 A549
TTTGTCATCCCAACGG-8 A549
TTTGTCATCTGGCGAC-8 A549
Name: cell_type, Length: 91205, dtype: category
Categories (1, object): ['A549']
print(adata_temp.obs['dose_val'])
cell_barcode
AAACCTGAGGCATGTG-1 1+1
AAACCTGAGGCCCTTG-1 1+1
AAACCTGCACGAAGCA-1 1
AAACCTGCAGACGTAG-1 1+1
AAACCTGCAGCCTTGG-1 1+1
...
TTTGTCAGTCATGCAT-8 1+1
TTTGTCATCAGTACGT-8 1+1
TTTGTCATCCACTCCA-8 1+1
TTTGTCATCCCAACGG-8 1+1
TTTGTCATCTGGCGAC-8 1+1
Name: dose_val, Length: 91205, dtype: category
Categories (2, object): ['1', '1+1']
print(adata_temp.obs['control'])
cell_barcode
AAACCTGAGGCATGTG-1 0
AAACCTGAGGCCCTTG-1 0
AAACCTGCACGAAGCA-1 1
AAACCTGCAGACGTAG-1 0
AAACCTGCAGCCTTGG-1 0
..
TTTGTCAGTCATGCAT-8 0
TTTGTCATCAGTACGT-8 0
TTTGTCATCCACTCCA-8 0
TTTGTCATCCCAACGG-8 0
TTTGTCATCTGGCGAC-8 0
Name: control, Length: 91205, dtype: int64
print(adata_temp.obs['condition_name'])
cell_barcode
AAACCTGAGGCATGTG-1 A549_TSC22D1+ctrl_1+1
AAACCTGAGGCCCTTG-1 A549_KLF1+MAP2K6_1+1
AAACCTGCACGAAGCA-1 A549_ctrl_1
AAACCTGCAGACGTAG-1 A549_CEBPE+RUNX1T1_1+1
AAACCTGCAGCCTTGG-1 A549_MAML2+ctrl_1+1
...
TTTGTCAGTCATGCAT-8 A549_RHOXF2BB+SET_1+1
TTTGTCATCAGTACGT-8 A549_FOXA3+ctrl_1+1
TTTGTCATCCACTCCA-8 A549_CELF2+ctrl_1+1
TTTGTCATCCCAACGG-8 A549_BCORL1+ctrl_1+1
TTTGTCATCTGGCGAC-8 A549_MAP4K3+ctrl_1+1
Name: condition_name, Length: 91205, dtype: category
Categories (284, object): ['A549_AHR+FEV_1+1', 'A549_AHR+KLF1_1+1', 'A549_AHR+ctrl_1+1', 'A549_ARID1A+ctrl_1+1', ..., 'A549_ctrl+UBASH3B_1+1', 'A549_ctrl+ZBTB1_1+1', 'A549_ctrl+ZBTB25_1+1', 'A549_ctrl_1']
print(adata_temp.var['gene_name'])
gene_id
ENSG00000239945 RP11-34P13.8
ENSG00000223764 RP11-54O7.3
ENSG00000187634 SAMD11
ENSG00000187642 PERM1
ENSG00000188290 HES4
...
ENSG00000198786 MT-ND5
ENSG00000198695 MT-ND6
ENSG00000198727 MT-CYB
ENSG00000273554 AC136616.1
ENSG00000278633 AC023491.2
Name: gene_name, Length: 5045, dtype: category
Categories (5045, object): ['A2M', 'A2M-AS1', 'A2ML1', 'AAK1', ..., 'ZSWIM1', 'ZSWIM4', 'ZSWIM8', 'ZYX']
print(adata_temp.uns['non_dropout_gene_idx'])
'A549_AHR+FEV_1+1': array([ 0, 2, 3, ..., 5042, 5043, 5044], dtype=int64), 'A549_AHR+KLF1_1+1': array([ 0, 1, 2, ..., 5042, 5043, 5044], dtype=int64),......
对于dixit数据集
pert_data.adata
View of AnnData object with n_obs × n_vars = 43401 × 5012
obs: 'condition', 'cell_type', 'dose_val', 'control', 'condition_name'
var: 'gene_name'
uns: 'non_dropout_gene_idx', 'non_zeros_gene_idx', 'rank_genes_groups_cov_all', 'top_non_dropout_de_20', 'top_non_zero_de_20'
layers: 'counts'
基本格式与Norman一致,但是它没有双扰动数据
print(pert_data.adata.obs[~pert_data.adata.obs['condition'].str.contains('ctrl', na=False)])
Empty DataFrame
Columns: [condition, cell_type, dose_val, control, condition_name]
Index: []
对于Adamson数据集
View of AnnData object with n_obs × n_vars = 65899 × 5060
obs: 'condition', 'cell_type', 'dose_val', 'control', 'condition_name'
var: 'gene_name'
uns: 'non_dropout_gene_idx', 'non_zeros_gene_idx', 'rank_genes_groups_cov_all', 'top_non_dropout_de_20', 'top_non_zero_de_20'
这个数据集和dixit数据集都是K562细胞
也没有双扰动
对于replogle_k562_essential数据集
View of AnnData object with n_obs × n_vars = 162246 × 5000
obs: 'condition', 'cell_type', 'cov_drug_dose_name', 'dose_val', 'control', 'condition_name'
var: 'gene_name', 'chr', 'start', 'end', 'class', 'strand', 'length', 'in_matrix', 'mean', 'std', 'cv', 'fano', 'highly_variable', 'means', 'dispersions', 'dispersions_norm'
uns: 'hvg', 'non_dropout_gene_idx', 'non_zeros_gene_idx', 'rank_genes_groups_cov_all', 'top_non_dropout_de_20', 'top_non_zero_de_20'
这个数据集也是k562类型的细胞
print(pert_data.adata.obs['cov_drug_dose_name'])
cell_barcode
AAACCCAAGAAGCCAC-34 K562_UBL5+ctrl_1+1
AAAGGATTCTCTCGAC-42 K562_UBL5+ctrl_1+1
AACGGGAGTAATGATG-25 K562_UBL5+ctrl_1+1
AAGAACAAGCTAGATA-35 K562_UBL5+ctrl_1+1
AAGACTCTCTATTGTC-33 K562_UBL5+ctrl_1+1
...
TTATTGCCACGTGAGA-26 K562_RPS2+ctrl_1+1
TTGTTGTTCATGGATC-47 K562_RPS2+ctrl_1+1
TTTCACATCTCTTCAA-41 K562_RPS2+ctrl_1+1
TTTCAGTGTAGAGTTA-18 K562_RPS2+ctrl_1+1
TTTCCTCTCGTTCGCT-42 K562_RPS2+ctrl_1+1
Name: cov_drug_dose_name, Length: 162246, dtype: category
Categories (1088, object): ['K562_AAMP+ctrl_1+1', 'K562_AARS+ctrl_1+1', 'K562_AATF+ctrl_1+1', 'K562_ABCB7+ctrl_1+1', ..., 'K562_ZNF574+ctrl_1+1', 'K562_ZNHIT6+ctrl_1+1', 'K562_ZNRD1+ctrl_1+1', 'K562_ctrl_1']
print(pert_data.adata.var['class'])
gene_id
ENSG00000237491 gene_version10
ENSG00000188290 gene_version10
ENSG00000187608 gene_version10
ENSG00000176022 gene_version7
ENSG00000131584 gene_version19
...
ENSG00000198695 gene_version2
ENSG00000278704 gene_version1
ENSG00000274847 gene_version1
ENSG00000278384 gene_version1
ENSG00000271254 gene_version6
Name: class, Length: 5000, dtype: category
Categories (26, object): ['gene_version1', 'gene_version2', 'gene_version3', 'gene_version4', ..., 'gene_version23', 'gene_version24', 'gene_version25', 'gene_version27']
print(pert_data.adata.var['strand'])
gene_id
ENSG00000237491 +
ENSG00000188290 -
ENSG00000187608 +
ENSG00000176022 +
ENSG00000131584 -
..
ENSG00000198695 -
ENSG00000278704 -
ENSG00000274847 -
ENSG00000278384 -
ENSG00000271254 -
Name: strand, Length: 5000, dtype: category
Categories (2, object): ['+', '-']
对于replogle_rpe1_essential数据集
View of AnnData object with n_obs × n_vars = 161423 × 5000
obs: 'condition', 'cell_type', 'cov_drug_dose_name', 'dose_val', 'control', 'condition_name'
var: 'gene_name', 'chr', 'start', 'end', 'class', 'strand', 'length', 'in_matrix', 'mean', 'std', 'cv', 'fano', 'highly_variable', 'means', 'dispersions', 'dispersions_norm'
uns: 'hvg', 'non_dropout_gene_idx', 'non_zeros_gene_idx', 'rank_genes_groups_cov_all', 'top_non_dropout_de_20', 'top_non_zero_de_20'
代码
对于seen数据的划分
def get_simulation_split(self,pert_list,trian_gene_set_size=0.85,combo_seen2_train_frac = 0.85,
seed = 1 , test_set_perts = None , only_test_set_perts = False ) :
"""
pert_list -> unqiue_perts
这里的unique_perts代表了不重复的扰动
"""
unique_pert_genes = self.get_genes_from_perts(pert_list)
"""
这里的unique_pert_genes代表了不重复扰动中不重复的基因
"""
pert_train = []
pert_test = []
np.random.seed(seed=seed)
if only_test_set_perts and ( test_set_perts is not None) :
ood_genes = np.array(test_set_perts)
train_gene_candidates = np.setdiff1d(unique_pert_genes,ood_genes)
else:
train_gene_candidates = np.random.choice ( unique_pert_genes,
int(len(unique_pert_genes) * trian_gene_set_size ) , replace = False)
if test_set_perts is not None:
num_overlap = len(np.intersect1d(train_gene_candidates,test_set_perts))
train_gene_candidates = train_gene_candidates[~np.isin(train_gene_candidates,test_set_perts)]
ood_genes_exclude_test_set = np.setdiff1d(unique_pert_genes,np.union1d(train_gene_candidates,test_set_perts))
train_set_addition = np.random.choice(ood_genes_exclude_test_set,num_overlap,replace=False)
train_gene_candidates = np.concatenate((train_gene_candidates,train_set_addition))
ood_genes = np.setdiff1d(unique_pert_genes,train_gene_candidates)
pert_single_train = self.get_perts_from_genes(train_gene_candidates,pert_list,'single')
## 这里的pert_combo包含了只要你的基因有一个在PertList里面,就从Pertlist把这个Pert提取出来
pert_combo = self.get_perts_from_genes(train_gene_candidates,pert_list,'combo')
pert_train.extend(pert_single_train)
## the combo set with one of them in OOD
combo_seen1 = [x for x in pert_combo if len([t for t in x.split('+') if
t in train_gene_candidates]) == 1]
pert_test.extend(combo_seen1)
pert_combo = np.setdiff1d(pert_combo,combo_seen1)
np.random.seed(seed=seed)
pert_combo_train = np.random.choice(pert_combo,int(len(pert_combo)*combo_seen2_train_frac),replace=False)
combo_seen2 = np.setdiff1d(pert_combo,pert_combo_train).tolist()
pert_test.extend(combo_seen2)
pert_train.extend(pert_combo_train)
## unseen single
unseen_single = self.get_perts_from_genes(ood_genes,pert_list,"single")
combo_ood = self.get_perts_from_genes(ood_genes,pert_list,'combo')
pert_test.extend(unseen_single)
combo_seen0 = [x for x in combo_ood if len([t for t in x.split('+') if t in train_gene_candidates]) == 0]
pert_test.extend(combo_seen0)
assert len(combo_seen0) + len(combo_seen1) + len(unseen_single) + len(pert_train) + len(combo_seen2) == len(pert_list)
return pert_train,pert_test,{'combo_seen0': combo_seen0,
'combo_seen1': combo_seen1,
'combo_seen2': combo_seen2,
'unseen_single': unseen_single}
#上面这个代码的combo_seen2并不能保证它的基因是都看过的